Not only in information technology: restart also works in chemical simulations
The innovative way of expediting chemical simulations: resetting them
A new study from Tel Aviv University discovered that a known practice in information technology can also be applied to chemistry. Researchers found that to enhance the sampling in chemical simulations, all you need to do is stop and restart.

(Left to right): Prof. Shlomi Reuveni, Ph.D. student Ofir Blumer & Dr. Barak Hirshberg.
Courtesy of Tel Aviv University
The research was led by Ph.D. student Ofir Blumer, in collaboration with Professor Shlomi Reuveni and Dr. Barak Hirshberg from the Sackler School of Chemistry at Tel Aviv University. The study was published in the journal Nature Communications.
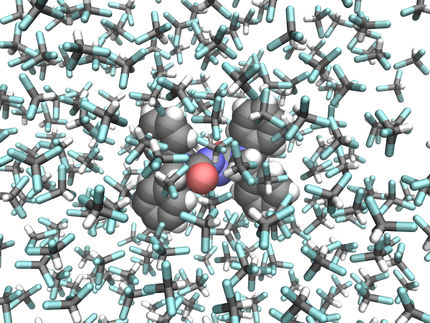
The researchers explain that molecular dynamics simulations are like a virtual microscope. They track the motion of all atoms in chemical, physical, and biological systems such as proteins, liquids, and crystals. They provide insights into various processes and have different technological applications, including drug design. However, these simulations are limited to processes slower than one-millionth of a second, and thus cannot describe slower processes such as protein folding and crystal nucleation. This limitation, known as the timescale problem, is a great challenge in the field.
Ph.D. student Ofir Blumer: “In our new study we show that the timescale problem can be overcome by stochastic resetting of the simulations. It seems counterintuitive at first glance – how can the simulations end faster when restarted? Yet, it turns out that reaction times vary considerably between simulations. In some simulations, reactions occur rapidly, but other simulations get lost in intermediate states for long periods. Resetting prevents the simulations from getting stuck in such intermediates and shortens the average simulation time.”
The researchers also combined stochastic resetting with Metadynamics, a popular method to expedite the simulations of slow chemical processes. The combination allows greater acceleration than either method separately. Moreover, Metadynamics relies on prior knowledge: the reaction coordinates must be known to expedite the simulation. The combination of Metadynamics with resetting reduces the dependency on prior knowledge significantly, saving time for practitioners of the method. Finally, the researchers showed that the combination provides more accurate predictions of the rate of slow processes. The combined method was used to enhance simulations of a protein folding in water successfully and it is expected to be applied to more systems in the future.
Original publication
Other news from the department science

Get the chemical industry in your inbox
By submitting this form you agree that LUMITOS AG will send you the newsletter(s) selected above by email. Your data will not be passed on to third parties. Your data will be stored and processed in accordance with our data protection regulations. LUMITOS may contact you by email for the purpose of advertising or market and opinion surveys. You can revoke your consent at any time without giving reasons to LUMITOS AG, Ernst-Augustin-Str. 2, 12489 Berlin, Germany or by e-mail at revoke@lumitos.com with effect for the future. In addition, each email contains a link to unsubscribe from the corresponding newsletter.